S3: Functional Initialization & Crop
Step Code: S3_func_init_and_crop
Depends on: S2 (Anatomical Cord Reference), BIDS Functional Data
Required by: S4 (Motion Correction)
Purpose
S3 prepares raw functional data for motion correction and downstream processing. Functional MRI of the spinal cord suffers from physiological noise, variable field of view, and susceptibility artifacts. This step ensures a clean starting point by:
- ** stabilizing the time series** (dropping dummy volumes)
- Locating the cord directly in functional space
- Identifing and gating outlier frames to create a robust reference
- Cropping the FOV to the spinal cord to reduce computational load and improve registration
Algorithm Overview
The S3 step consists of three sequential subtasks:
| Subtask | Name | Description |
|---|---|---|
| S3.1 | Dummy Drop & Localization | Remove initial unstable frames, compute fast reference, localize cord |
| S3.2 | Outlier Gating | Compute DVARS/RefRMS metrics, identify bad frames, create robust reference |
| S3.3 | Crop & QC | Apply cord-focused crop, generate quality control reportlets |
S3.1: Dummy Drop & Localization
Rationale
MRI scanners require a few TRs to reach steady-state magnetization. These initial "dummy" volumes have inconsistent contrast and should be discarded. Subsequent processing requires an approximate location of the spinal cord to focus analysis.
Algorithm
- Dummy Volume Drop - Remove first $N$ volumes (default: 4)
- Fast Reference - Compute median of all remaining frames
- Cord Localization - Segment cord on fast reference using
sct_deepseg
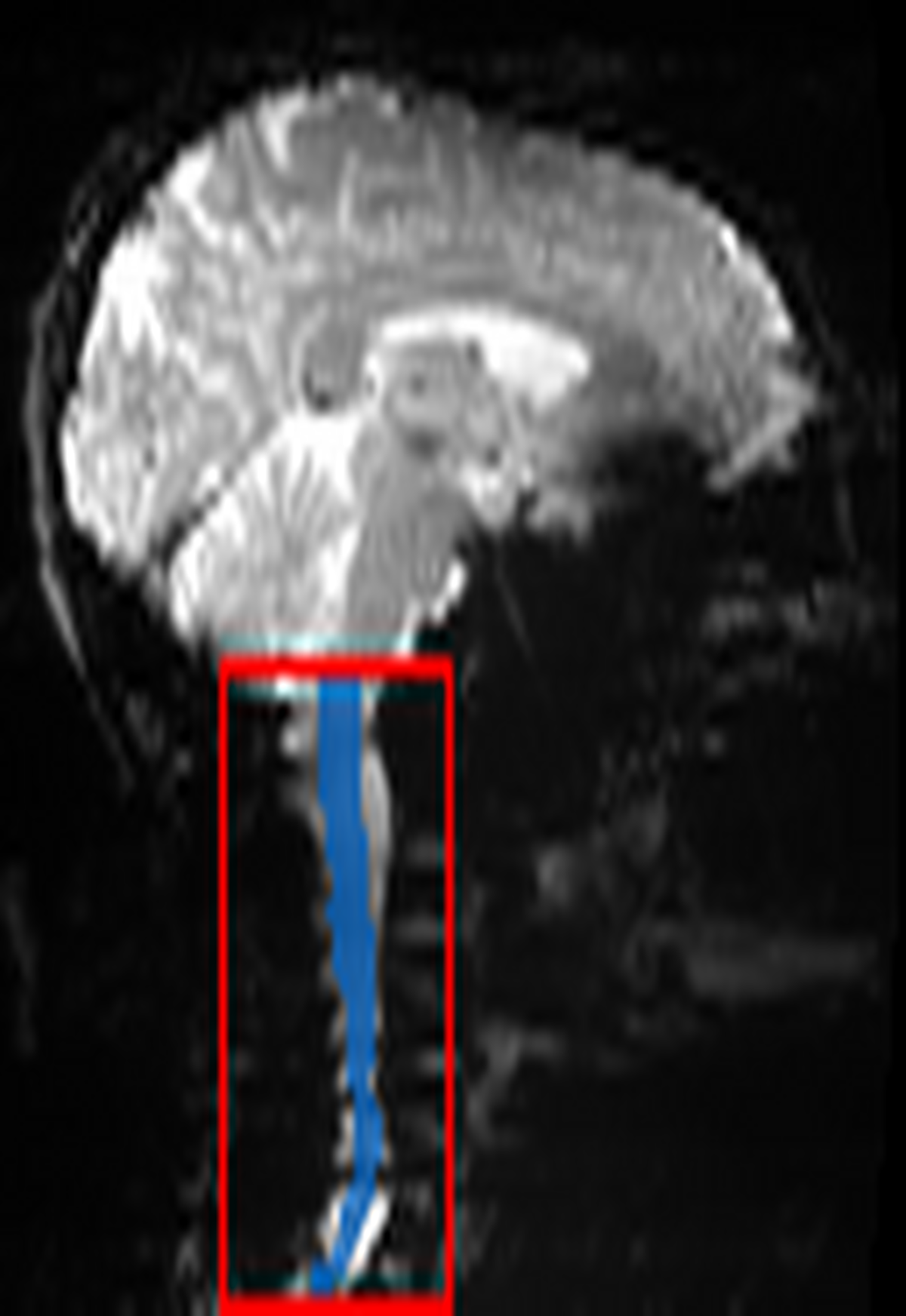
QC: Functional Localization
What to look for:
- ✅ Blue contour (cord mask) accurately follows the spinal cord in the functional image.
- ✅ Red box centers on the cord and includes sufficient margin.
- ❌ FAIL: Mask is empty or tracks non-cord structure (e.g., ghosting artifacts).
S3.2: Outlier Gating
Rationale
Subject motion and physiological noise can corrupt specific frames. Using a simple mean/median of all frames as a reference for motion correction can bake in these artifacts. Outlier gating ensures the reference image is constructed only from stable, high-quality frames.
Algorithm
- Metric Computation (within cord mask):
- DVARS: Derivative of variance (frame-to-frame change)
- RefRMS: Root mean square error vs. fast reference
- Outlier Detection:
- Threshold = $P_{75} + 1.5 \times IQR$
- Frame is outlier if DVARS or RefRMS exceeds threshold
- Robust Reference:
- Compute median of only non-outlier frames
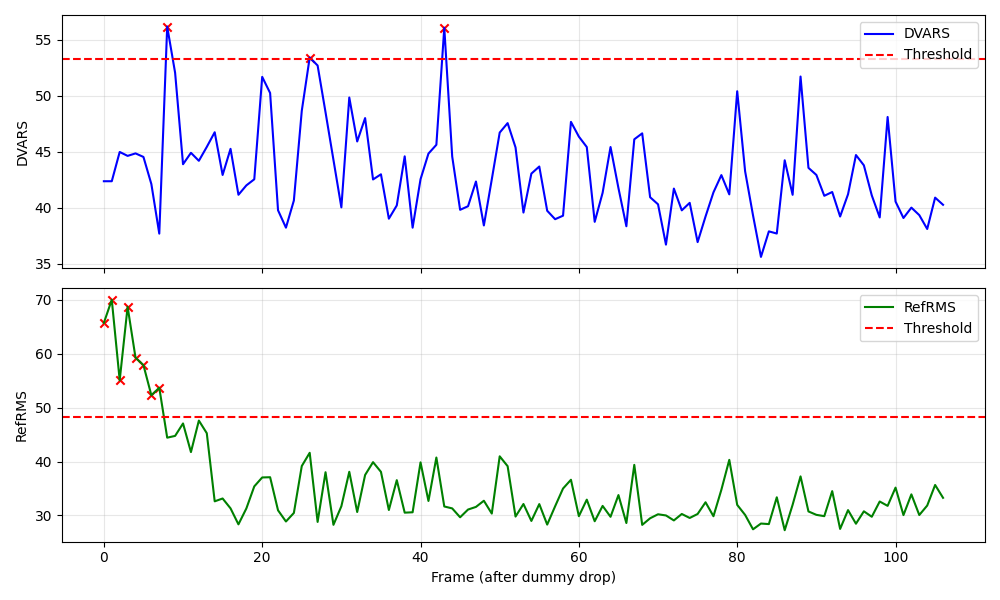
QC: Frame Metrics
What to look for:
- ✅ Metrics (blue/orange lines) vary but stay mostly below thresholds (dashed lines).
- ✅ Outliers (if any) correspond to visible spikes in the plot.
- ❌ FAIL: Majority of frames exceed thresholds (suggests severe artifact or bad mask).
- ❌ FAIL: Periodic spikes aligned with respiration (if TR is aliased with breathing).
S3.3: Crop & QC
Rationale
Processing the full field-of-view (FOV) is unnecessary and computationally expensive. A tight crop around the spinal cord improves the robustness of subsequent motion correction (S4) by excluding moving structures like the throat and chest.
Algorithm
- Cylindrical Crop Mask - Create dynamic mask around centerline
- Apply Crop - Cut 4D dataset to crop mask extent
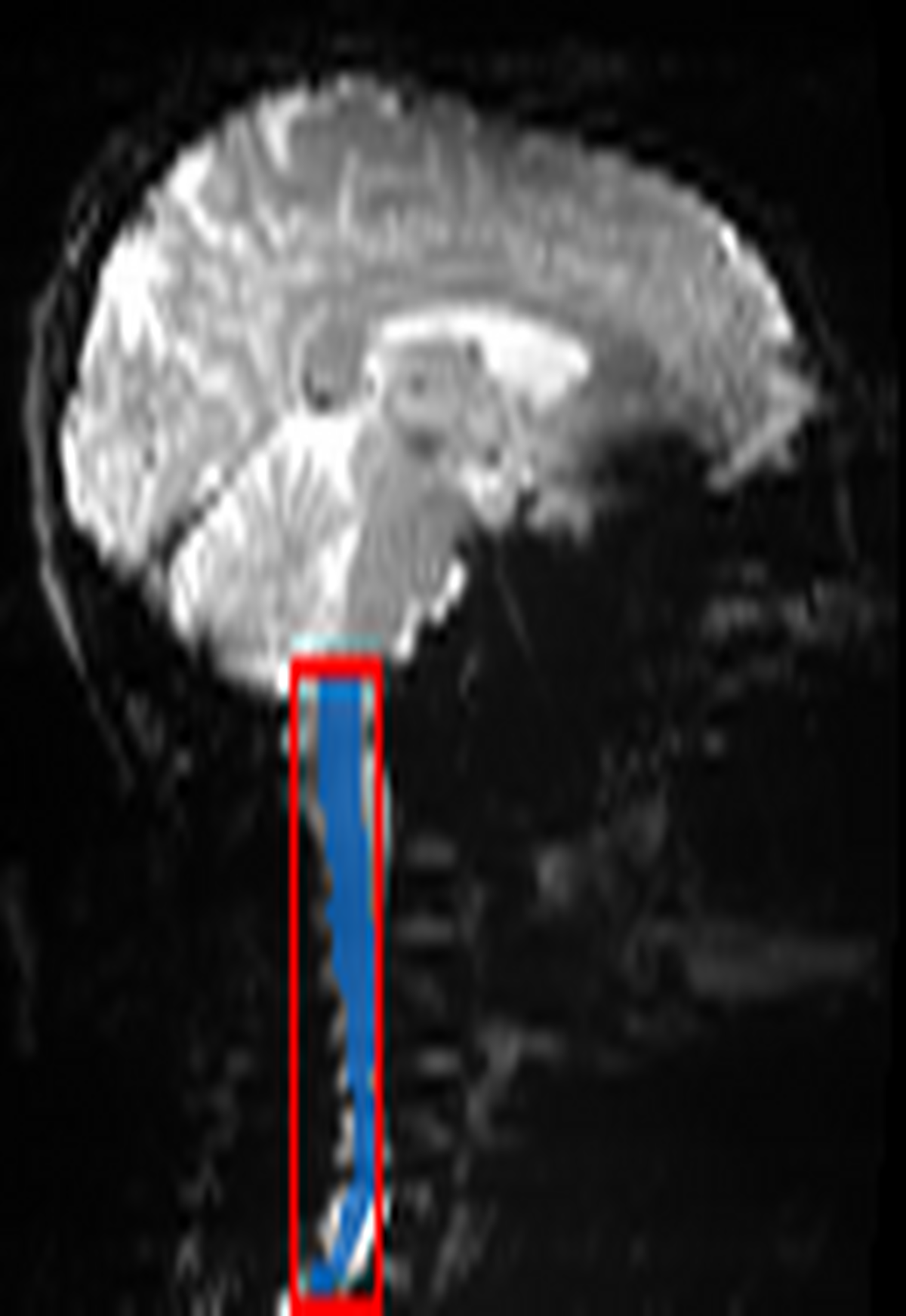
QC: Crop Box Sagittal
What to look for:
- ✅ Red box captures the entire spinal cord segment of interest.
- ✅ Box is not excessively large (should be ~40mm wide).
- ❌ FAIL: Cord leaves the crop box at top or bottom.
QC: Functional Reference Montage
What to look for:
- ✅ Clear definition of spinal cord against CSF (bright CSF, dark cord in T2*-weighted).
- ✅ No severe blurring (indicates robust reference construction worked).
- ❌ FAIL: "Ghosting" or double-images of the cord.
Outputs
Derivatives
derivatives/spinalfmriprep/{dataset}/sub-{id}/func/
├── sub-{id}_task-{task}_desc-funcref.nii.gz # Robust reference
├── sub-{id}_task-{task}_desc-funccrop_bold.nii.gz # Cropped 4D data
├── sub-{id}_task-{task}_desc-funccrop_mask.nii.gz # Binary crop mask
├── sub-{id}_task-{task}_desc-confounds_timeseries.tsv # Frame metrics (DVARS, etc.)
└── sub-{id}_task-{task}_desc-outliers.json # List of outlier indices
derivatives/spinalfmriprep/{dataset}/sub-{id}/figures/
├── sub-{id}_..._desc-S3_func_localization_crop_box_sagittal.png
├── sub-{id}_..._desc-S3_frame_metrics.png
├── sub-{id}_..._desc-S3_crop_box_sagittal.png
└── sub-{id}_..._desc-S3_funcref_montage.png
CLI Usage
# Run S3 for a single dataset
poetry run spinalfmriprep run S3_func_init_and_crop \
--dataset-key <KEY> \
--datasets-local config/datasets_local.yaml \
--out work/wf_reg_001
# Run S3 with parallel workers
poetry run spinalfmriprep run S3_func_init_and_crop \
--scope reg \
--out work/wf_reg_001 \
--batch-workers 8
QC Status Logic
Automated status determination based on metrics:
status = FAIL if:
- Cord localization fails (no mask generated)
- Outlier fraction > 50% (data too noisy)
- Fewer than 10 good frames remain
- Crop produces < 10 Z slices
status = WARN if:
- Outlier fraction > 30%
status = PASS otherwise
References
- SCT DeepSeg: Gros et al. NeuroImage 184:901-915 (2019). DOI
- DVARS: Power et al. NeuroImage 59(3):2142-2154 (2012). DOI
Last updated: February 2026